Update 2 August 2009: I have changed the discussion regarding the SG1 keyword. New text is in italics.I have planned some posts in which some details of the GAMESS input will become important, so this post is about the basic anatomy of a GAMESS input file, and some non-default settings that I use all the time.
Here is an input file for a single point energy calculation at the B3LYP/6-31G(d) level of theory for the H
2 molecule:
$contrl runtyp=energy dfttyp=b3lyp $end
$basis gbasis=n31 ngauss=6 ndfunc=1 $end
$data
Title line: you can write anything here
c1
H 1.0 0.0 0.0 0.0
H 1.0 0.753 0.0 0.0
$end• The order of the groups doesn’t matter, so you can put the $basis group below the $data group.
• You can have several groups of the same kind (e.g. two $contrl groups). If you have the same keyword in both groups, it is the value in the last group that counts. For example,
$contrl runtyp=energy dfttyp=b3lyp $end
$contrl runtyp=optimize $end is perfectly legal, and will result in an geometry optimization.
• The order of the keywords within a group doesn’t matter.
$contrl runtyp=energy dfttyp=b3lyp $end is the same as
$contrl dfttyp=b3lyp runtyp=energy $end• Note that there is a space in front of any line starting with a $ sign. This is important and if you leave it out GAMESS will not read that line. Note you can use this to “uncomment” a line by adding, for example, a ! For example, this
$contrl runtyp=energy dfttyp=b3lyp $end
!$contrl runtyp=optimize $end will result in a single point energy calculation
• You can use capital letters if you want.
• For parameters that can be true for false, you can use either “.true.” or “.t.” and “.false.” or “.f.” (there's an example below).
• A line can only be 80 characters long. Anything after that is ignored. It is perfectly legal to split a line, for example:
$contrl runtyp=energy
dfttyp=b3lyp $endSome useful optionsAlmost all of the parameters have default (but there is no default basis set and no defaults in $data). For example, the default runtyp is energy, so
$contrl dfttyp=b3lyp $end
will result in a single point energy evaluation. If $contrl is missing entirely, GAMESS will compute a single point energy at the RHF level.
There is a few defaults that I almost always change
Single point energy calculations (not AM1 or PM3)
$scf dirscf=.t. $endThis tells GAMESS to recompute the integrals at each SCF step rather than reading them from hard disk. Most computers can recompute a number faster than they can find it on the hard disk.
DFT single point energy calculations$dft sg1=.t. $endComputing the DFT energy requires a numerical integration over a grid. The default grid is very fine (has a lot of points). Standard Grid 1 (sg1) is adequate for most purposes and much faster.
Note that this leads to inaccurate gradients and should not be used for geometry optimizations.DFT, AM1, and PM3 frequency calculations $force nvib=2 $endDFT and semiempirical Hessians (second energy derivatives) are computed numerically, by displacing each atom in the
x,
y, and
z direction and computing the first derivative. This is usually not accurate enough. nvib=2 leads to a displacement in both the positive and negative
x,
y, and
z direction resulting in more accurate frequencies.
Geometry optimizations $statpt opttol=0.0005 nstep=50 $end
$contrl nzvar=1 $end
$zmat dlc=.t. auto=.t. $endI think the default criterion for geometry convergence (0.0001) is too strict, and the default number of steps (2o) is too small. So I usually use 0.0005 and 50, respectively. Optimizing in internal coordinates instead of Cartesian coordinates (default) usually speeds up convergence, because the former are "more quadratic". nzvar=1 tells GAMESS to use internal coordinates and dlc and auto tells GAMESS to automatically generate delocalized internal coordinates.
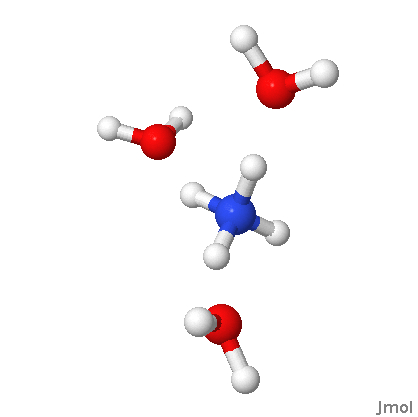
Unfortunately, the DLC implementation in GAMESS only looks for covalent bonds, not H-bonds and such. So you have to specify these if you have specify non-covalently bonded molecules in the input file. For example, in the case of water dimer (see figure for numbering), you do it like this
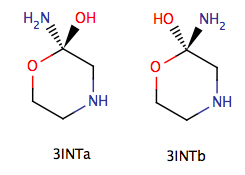
$zmat nonvdw(1)=1,4 $endNote that you don't have to specify the H-bond in HO-CH2-CH2-OH, since the OH groups are connected by covalent bonds.

So, an input file for a geometry optimization of the water dimer would look like what I show below. That might seem a lot to type, but the trick is of course to type is once, store it somewhere handy, and copy it into new files.
$contrl runtyp=optimize dfttyp=b3lyp $end
$basis gbasis=n31 ngauss=6 ndfunc=1 $end
$scf dirscf =.t. $end
$statpt opttol=0.0005 nstep=50 $end $contrl nzvar=1 $end
$zmat dlc=.t. auto=.t. nonvdw(1)=1,4 $end
$DATA
Title
C1
O 8.0 -4.42170 -0.11056 -2.11879
H 1.0 -3.53334 -0.03874 -1.70405
H 1.0 -4.75257 0.79398 -2.01675
O 8.0 -2.06207 0.35222 -0.78539
H 1.0 -1.13417 0.41136 -1.06880
H 1.0 -1.98676 0.07447 0.14406
$END